APPLICATIONS OF TECHNOLOGY:
- Screening platforms for pharmaceuticals
- Biological production of biofuels
- Tissue engineering
- Cell-based biosensing
ADVANTAGES:
- Faster and more biocompatible than existing methods
- Works for virtually all mammalian, plant, bacterial, and fungal cells
- Minimal to no effect on intracellular signaling pathways
- Enables patterning of multiple cell types in complex arrangements
- Applicable to diverse surface types (e.g., slides, electrodes)
- Reusable substrate
ABSTRACT:
Scientists at Berkeley Lab have developed an efficient technology for attaching cells to surfaces (substrates) in predetermined patterns. The method covalently links DNA strands to cell membranes. The cells then adhere to a surface that has been coated with a pattern of complementary oligonucleotides. Berkeley Lab scientists tested this technology by capturing cells from various biological kingdoms onto glass slides, atomic force microscope (AFM) probes, and metal electrodes used to measure cellular activities. They were able to attach different cell types to a single surface or to one another in solution.
In contrast with existing methods for patterning cells on a surface, the Berkeley Lab technology can be used with both adherent and nonadherent cells (e.g., blood and immune cells) from multiple species. It requires much less time (1 hour vs. 3 days) because it efficiently captures a much higher percentage of the cells exposed to the surface, circumventing the need for prolonged culture to grow the cells to confluence. The new method does not trigger intracellular pathways and differentiation, as occurs when integrins are bound. In addition, the surface can be reused by heating it enough to melt the DNA and release the cells to allow for the application of new ones. The technology can also capture an individual cell to make a single-cell microarray. The Berkeley Lab researchers believe that the strength of the different connections between cells and surfaces could potentially be controlled by selecting different lengths of complementary DNA strands.
In general, patterning cells on surfaces has provided a platform for studying individual cell function, cell-cell interactions, differentiation, and the engineering of new tissues (e.g., assembling muscle tissue in vitro). However, existing methods for patterning cells on a surface may be used only with integrin-bearing, adherent cells from certain species, and the methods often alter the biology of the cells. The Berkeley Lab technology overcomes these significant limitations.
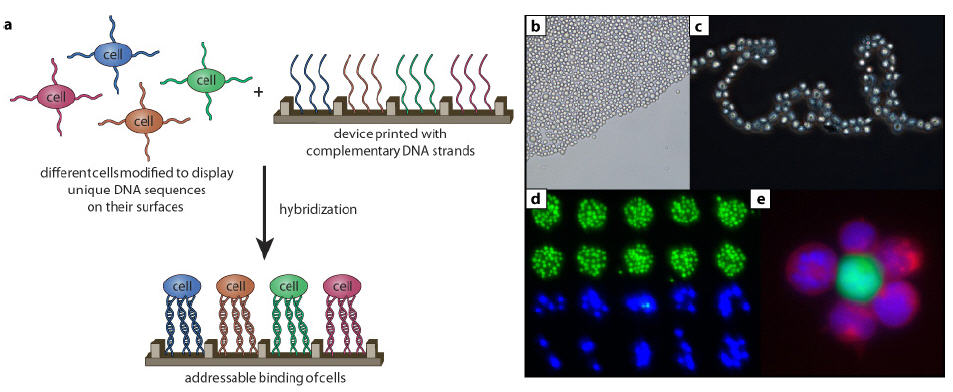
(a) Schematic for attaching living cells to surfaces through DNA hybridization.
(b) DNA-coated, cultured cancer (Jurkat) cells form closely packed arrays within minutes of exposure to surfaces bearing the DNA sequence complement.
(c) Jurkat cells can be arranged in complex patterns by printing “trails” of DNA that are 20 micrometers wide (about the width of one-cell).
(d) Multiple cell types can be captured from a single solution: Jurkat cells appear green and MDA breast cancer cells appear blue.
(e) Application of the method to cells in solution that have been labeled with different cytoplasmic dyes.
DEVELOPMENT STAGE: Bench scale prototype.
STATUS: Published Patent Application WO2010118235 available at www.wipo.int. Available for licensing or collaborative research.
SEE THESE OTHER BERKELEY LAB TECHNOLOGIES IN THIS FIELD:
A Cell Injection System Using Carbon Nanotubes, IB-2323, IB-2333
Membrane-Derivatized Colloids for the Detection and Analysis of Cell Membrane Binding Interaction, IB-1919
REFERENCE NUMBER: JIB-2703
